The
recent level one class (MBB165) on the influence of temperature and pH
on the enzymatic reaction catalysed by aryl sulfatase made me think
about explanations of classic "physical chemistry" effects on proteins
in general. School Biology and Chemistry classes introduce students to
the idea of pH and temperature optima for enzymes. In the case of
temperature, explanations are pretty straight forward: as the
temperature of a reaction increases, so too does the rate. You will
recall that the reason for this in a typical uncatalysed reaction is not
simply an increase in the number of molecular collisions, but a shift
in the proportion of molecules that
have crossed the "activation energy" threshold. Hence there is an
approximate doubling of reaction rate for every 10 degree increase in
temperature. However, when an enzyme is present, the increase is a
consequence )or function) not only of the activation effect, but is also a function of the intrinsic thermostability of the enzyme. So, enzymes exhibit temperature optima as a consequence of reaction kinetics and protein stability factors.
The
recent level one class (MBB165) on the influence of temperature and pH
on the enzymatic reaction catalysed by aryl sulfatase made me think
about explanations of classic "physical chemistry" effects on proteins
in general. School Biology and Chemistry classes introduce students to
the idea of pH and temperature optima for enzymes. In the case of
temperature, explanations are pretty straight forward: as the
temperature of a reaction increases, so too does the rate. You will
recall that the reason for this in a typical uncatalysed reaction is not
simply an increase in the number of molecular collisions, but a shift
in the proportion of molecules that
have crossed the "activation energy" threshold. Hence there is an
approximate doubling of reaction rate for every 10 degree increase in
temperature. However, when an enzyme is present, the increase is a
consequence )or function) not only of the activation effect, but is also a function of the intrinsic thermostability of the enzyme. So, enzymes exhibit temperature optima as a consequence of reaction kinetics and protein stability factors. How might you design experiments to dissect out these two components?
Can you think of an important enzyme catalysed reaction (used by thousands of labs every day) which relies heavily on thermal control?
The pH dependence of a reaction turns out to be a little more complex and you need to first consider the pH dependence of the side chains of the amino acids found in proteins. I wont discuss it here, but the same would be true for the bases found in RNA, in the case of catalytic RNA molecules, or ribozymes. That's for another time.
Amino acids and their pKas. All of the common amino acids except glycine are chiral (have a handedness). The major form found in Nature are the S-amino acid, or L-amino acids . [The mirror image of each amino acid can be found in Nature, although they are less common.] In the Table below, the third, fourth and fifth columns give pKa values of groups: -NH3 refers to the protonated alpha-amino group, the CO2H refers to the carboxylic acid group on the alpha-carbon, and the side chain is only relevant for a few amino acids. And this is a very important point. The impact of pH on protein structure and function is therefore limited to the acidic class (Glu and Asp), the basic pair; Lys and Arg and importantly, the amino acid histidine which has a "special" place in proteins, since its pKa lies in the neutral or physiological range. In all cases below, pKas refer to the free amino acid, and not the amino acid incorporated into a polypeptide or protein chain. Usually, we assume that not much happens to the pKa of the side chain on incorporation. While this is not strictly true, it is a good starting point, until the value can be measured independently. Polar side chains such as Ser, Thr, Tyr and Cys are important as Hydrogen bond acceptors, but in order to promote their reactivity, they generally need to be "influenced" by adjacent (in spatial terms) side chains, cofactors and in particular metal ions. Perhaps the best example of the key role of a Ser residue and the sphere of influence of the active site residues is given by the Serine Proteases.
Amino Acid
|
R
|
-NH3+
|
-CO2H
|
Side chain
|
pI
|
|---|---|---|---|---|---|
| Glycine, Gly | -H | 9.78 | 2.35 | 5.97 | |
| Alanine, Ala | -CH3 | 9.87 | 2.35 | 6.02 | |
| Valine, Val | -CH(CH3)2 | 9.74 | 2.29 | 5.97 | |
| Leucine, Leu | CH2CH(CH3)2 | 9.74 | 2.33 | 5.98 | |
| Isoleucine, Ile | CH(CH3)CH2CH3 | 9.76 | 2.32 | 6.02 | |
| Phenylalanine, Phe | 9.31 | 2.20 | 5.48 | ||
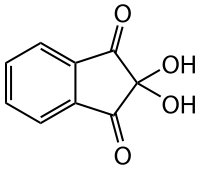
| Tryptophan, Trp |  |
9.41 | 2.46 | 5.88 | |
| Tyrosine, Tyr | 9.21 | 2.20 | 10.46 | 5.65 | |
| Histidine, His | 9.33 | 1.80 | 6.04* | 7.58 | |
| Serine, Ser | CH2OH | 9.21 | 2.19 | 5.68 | |
| Threonine, Thr | CH(CH3)-OH | 9.10 | 2.09 | 6.53 | |
| Methionine, Met | CH2CH2SCH3 | 9.28 | 2.13 | 5.75 | |
| Cysteine, Cys | CH2SH | 10.70 | 1.92 | 8.37 | 5.14 |
| Aspartic Acid, Asp | CH2CO2H | 9.90 | 1.99 | 3.90 | 2.87 |
| Glutamic Acid, Glu | CH2CH2CO2H | 9.47 | 2.10 | 4.07 | 3.22 |
| Asparagine, Asn | CH2CONH2 | 8.72 | 2.14 | 5.41 | |
| Glutamine, Gln | CH2CH2CONH2 | 9.13 | 2.17 | 5.65 | |
| Lysine, Lys | (CH2)4NH2 | 9.06 | 2.16 | 10.54* | 9.74 |
| Arginine, Arg | 8.99 | 1.82 | 12.48* | 10.76 | |
| Proline, Pro | 10.64 | 1.95 | 6.10 |
In order to explain the pH dependence of enzymes we need to two major factors: (i) folding and stability and (ii) orientation and reactivity of active site side chains. These topics will be covered in great detail during your degree, but here is a taster of things to come.
Protein folding and stability is determined by well established principles of Physical Chemistry combined with the unique chemistry of biological polypeptides. As Anfinsen has discussed: the primary structure of a protein carries sufficient information to direct its folding into a uniquely active conformation. [There are some emerging caveats to this principle, but they can come later]. There are some difficulties encountered in the experimental investigation of protein folding and so despite Anfinsen's insight and choice of the "well-behaved" nucleases, detailed investigations of protein folding really became possible with the advent of high resolution NMR spectrometry. The second barrier to understanding the mechanism of protein folding has become known as Levinthal's paradox. In short, for a given polypeptide chain comprising say 200 amino acid monomer units, each monomer unit has the freedom to sample many conformations (rotation about bonds etc), and the time taken for such a polypeptide to sample all possible structures is incompatible with "biological time". Clearly, the cell has a solution and in level 3 you will come across lowest free energy states and the "folding funnel" model for protein folding. Let's simplify things here and consider the following stages of folding of a polypeptide chain emerging from the ribosome or a round bottom flask in the lab. First, the hydrophobic effect drives the polypeptide chain into several more compact forms, the acquisition of secondary structure elements often then leads to the formation of a metastable state and finally, the lowest free energy form of the functional protein is attained, usually in less than 1ms. I think it is pretty clear that whilst many secondary structure interactions involve main chain hydrogen bond donors and acceptors, the later stage tuning and stabilisation of the final structure involves side chain interactions. As a consequence, the potential for a pH influence becomes inevitable.The pH of the solvent can influence the properties of the active site of an enzymes by perturbing the ionisation of key residues. As you will have noticed, the pH profile of an enzyme is often (but not always) bell shaped. The peak of the curve represents the optimum pH for activity, whilst the descending "shoulders" tend to reflect the combined influence of pH on protein stability and activity:at extremes of pH, most proteins tend to denature. An interesting couple of examples to consider from the literature are the gastric protease pepsin and the Krebs Cycle enzyme fumarase. Pepsin is a proteolytic enzyme that is usually found at acidic pHs: interestingly as the pH of the solvent is raised from 5 to 7, pepsin begins to denature. Not surprisingly, it is an acid protease and aspartic acid is found at its active site.
Can you think of an aspartic protease that made the headlines as a potential drug target?
Fumarase was characterised in a lovely early pH study by Frieden and Alberty: two well known enzymologists. In a paper published over 50 years ago, they demonstrated the insight that could be obtained from a careful analysis of the pH dependence of an enzymatic reaction. Such studies underpin the broad view that ionisation of two or more key residues at the active site of an enzyme, is responsible for the pH profile. If we look at the active site of aryl sulfatases, not the one from the snail, Helix pomatia (the source used in the lab classes),but from a related enzyme found in bacteria (for which there is a very high resolution structure), it is immediately clear that it is stuffed full of ionisable side chains: see below.
The other interesting feature is the calcium ion at the centre. Can you think of a plausible mechanism for how such a clustering of residues could promote the removal of the sulphate group from the aromatic ring?
In summary, pH has the potential to influence protein function through structural perturbations and by modifying the degree of ionisation of key active site side chains. As you can probably tell, methods of analysis are needed that identify the contribution of all amino acids in a protein during catalysis. Such methods will come, but for now we will continue to make educated guesses to supplement experimental data.